Columns have mixed types. Specify dtype option on import
Last updated: Apr 11, 2024
Reading time·5 min

# Table of Contents
- Columns have mixed types. Specify dtype option on import
- Silencing the warning by setting
dtypetoobject - Silencing the warning by setting low_memory to False
- Using the unicode data type instead
- Explicitly setting the engine to python
- Using converters to resolve the warning
# Columns have mixed types. Specify dtype option on import
The Pandas warning "Columns have mixed types. Specify dtype option on import or set low_memory=False." occurs when your CSV file contains columns that have mixed types and cannot be inferred reliably.
To resolve the issue, specify the data type of each column explicitly by
supplying the dtype argument.
For example, suppose you have the following employees.csv file.
first_name,last_name,date,salary Alice,Smith,01/21/1995,1500 Bobby,Hadz,04/14/1998,abc Carl,Lemon,06/11/1994,3000 Dean,Berry,06/11/1996,xyz
Notice that the salary column contains integers and strings.
The best way of solving the issue where pandas tries to guess the data type of
the column is to explicitly specify it.
import pandas as pd dtype = { 'first_name': str, 'last_name': str, 'date': str, 'salary': str } df = pd.read_csv( 'employees.csv', sep=',', encoding='utf-8', dtype=dtype ) # first_name last_name date salary # 0 Alice Smith 01/21/1995 1500 # 1 Bobby Hadz 04/14/1998 abc # 2 Carl Lemon 06/11/1994 3000 # 3 Dean Berry 06/11/1996 xyz print(df)
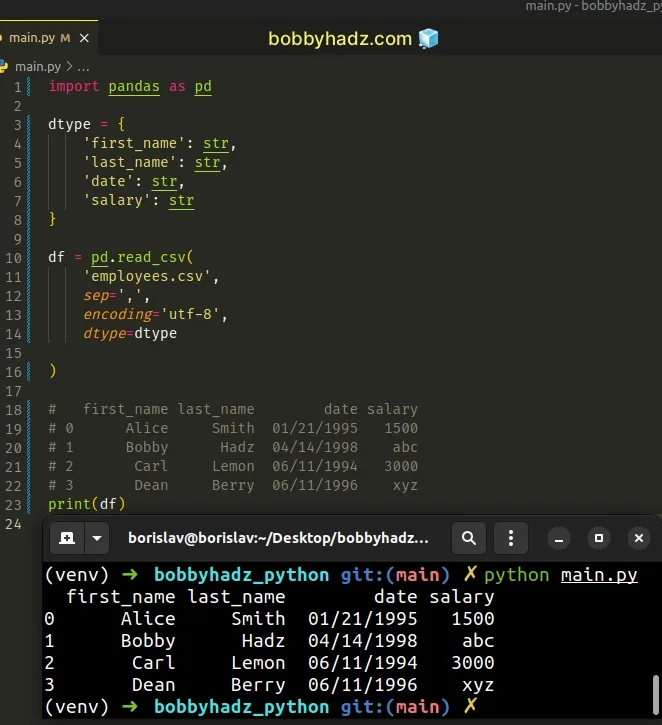
The dtype argument of the
pandas.read_csv() method
is a dictionary of column name -> type.
Here is another example.
import numpy as np dtype={'a': np.float64, 'b': np.int32, 'c': 'Int64'}
You can use the str or object types to preserve and not interpret the
dtype.
Pandas can only infer the data type of a column once the entire column is read.
In other words, pandas can't start parsing the data in the column until all of
it is read which is very inefficient.
employee.csv file with a salary column that only contains numbers, pandas can't know that the column only contains numbers until it has read all 5 million rows.Note that the salary column in the CSV file doesn't contain only integers.
first_name,last_name,date,salary Alice,Smith,01/21/1995,1500 Bobby,Hadz,04/14/1998,abc Carl,Lemon,06/11/1994,3000 Dean,Berry,06/11/1996,xyz
So if you try to set the dtype of the column to int, you'd get an error:
import pandas as pd dtype = { 'first_name': str, 'last_name': str, 'date': str, 'salary': int # 👈️ set dtype to int } # ⛔️ ValueError: invalid literal for int() with base 10: 'abc' df = pd.read_csv( 'employees.csv', sep=',', encoding='utf-8', dtype=dtype )
# Silencing the warning by setting dtype to object
If you just want to silence the warning, set the dtype of the column to
object.
import pandas as pd dtype = { 'first_name': str, 'last_name': str, 'date': str, 'salary': object } df = pd.read_csv( 'employees.csv', sep=',', encoding='utf-8', dtype=dtype ) # first_name last_name date salary # 0 Alice Smith 01/21/1995 1500 # 1 Bobby Hadz 04/14/1998 abc # 2 Carl Lemon 06/11/1994 3000 # 3 Dean Berry 06/11/1996 xyz print(df)
Setting the dtype of the column to object silences the warning but doesn't
make the read_csv() method more memory efficient.
# Silencing the warning by setting low_memory to False
You can also silence the warning by setting low_memory to False in the call
to pandas.read_csv().
import pandas as pd df = pd.read_csv( 'employees.csv', sep=',', encoding='utf-8', low_memory=False ) # first_name last_name date salary # 0 Alice Smith 01/21/1995 1500 # 1 Bobby Hadz 04/14/1998 abc # 2 Carl Lemon 06/11/1994 3000 # 3 Dean Berry 06/11/1996 xyz print(df)
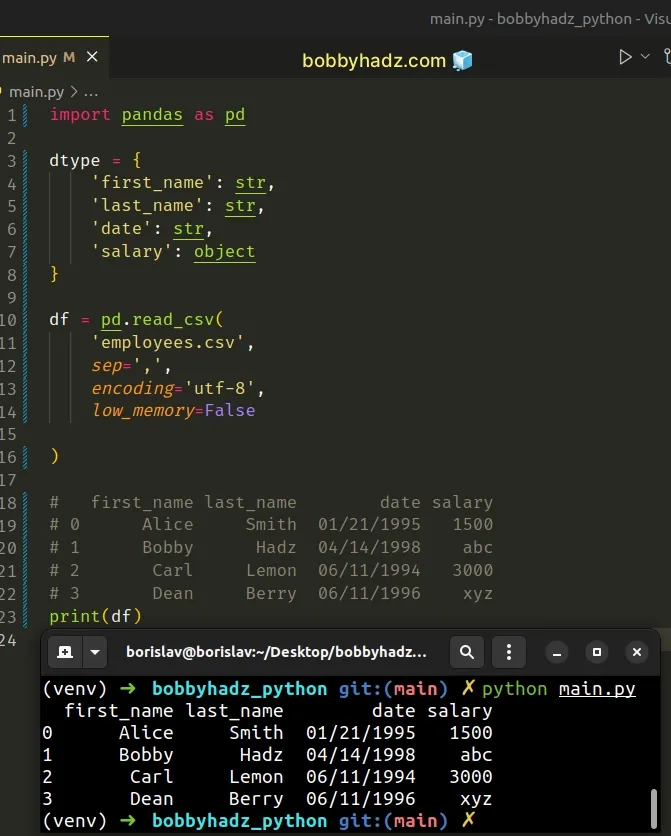
By default, the low_memory argument is set to True.
When the argument is set to True, the file is processed in chunks, resulting
in lower memory consumption while parsing, but possibly mixed type inference.
To ensure no mixed types, you can set the low_memory argument to False or
explicitly set the dtype parameter as we did in the previous subheading.
# Using the unicode data type instead
You can also silence the warning by using the unicode data type.
import pandas as pd import numpy as np df = pd.read_csv( 'employees.csv', sep=',', encoding='utf-8', index_col=False, dtype=np.dtype('unicode'), ) # first_name last_name date salary # 0 Alice Smith 01/21/1995 1500 # 1 Bobby Hadz 04/14/1998 abc # 2 Carl Lemon 06/11/1994 3000 # 3 Dean Berry 06/11/1996 xyz print(df)
Make sure you have the NumPy module installed by running the following command.
pip install numpy # or with pip3 pip3 install numpy
We set the data type to Unicode strings.
When index_col is set to False, pandas won't use the first column as the
index (e.g. when you have a malformed CSV file with delimiters at the end of
each line).
# Explicitly setting the engine to python
Another thing you can try is to set the engine argument to python when
calling pandas.read_csv.
import pandas as pd df = pd.read_csv( 'employees.csv', sep=',', encoding='utf-8', engine='python' ) # first_name last_name date salary # 0 Alice Smith 01/21/1995 1500 # 1 Bobby Hadz 04/14/1998 abc # 2 Carl Lemon 06/11/1994 3000 # 3 Dean Berry 06/11/1996 xyz print(df)
The engine argument is the parser engine that should be used.
The available engines are c, pyarrow and python.
The c and pyarrow engines are faster, however, the python engine is more
feature-complete.
# Using converters to resolve the warning
You can also use converters to resolve the warning.
Suppose you have the following employees.csv file.
first_name,last_name,date,salary Alice,Smith,01/21/1995,1500 Bobby,Hadz,04/14/1998,abc Carl,Lemon,06/11/1994,3000 Dean,Berry,06/11/1996,xyz
Notice that the salary column contains integers and strings.
Here is how we can use a converter to initialize the non-numeric salary values
to 0.
import pandas as pd import numpy as np def converter1(value): if not value: return 0 try: return np.float64(value) except Exception as _e: return np.float64(0) df = pd.read_csv( 'employees.csv', sep=',', encoding='utf-8', converters={'salary': converter1} ) # first_name last_name date salary # 0 Alice Smith 01/21/1995 1500.0 # 1 Bobby Hadz 04/14/1998 0.0 # 2 Carl Lemon 06/11/1994 3000.0 # 3 Dean Berry 06/11/1996 0.0 print(df)
The example only uses a converter for the salary column, however, you can
specify as many converters as necessary in the converters dictionary.
The converter1 function gets called with each field from the salary column.
If the supplied value is empty, we return 0.
The try block tries to convert the value to a NumPy float.
If an error is raised the except block runs where we return a 0 value.
# Additional Resources
You can learn more about the related topics by checking out the following tutorials:
- You are trying to merge on int64 and object columns [Fixed]
- Copy a column from one DataFrame to another in Pandas
- ValueError: cannot reindex on an axis with duplicate labels
- ValueError: Length mismatch: Expected axis has X elements, new values have Y elements
- ValueError: cannot reshape array of size X into shape Y
- Object arrays cannot be loaded when allow_pickle=False
- ValueError: Columns must be same length as key [Solved]
- ValueError: DataFrame constructor not properly called [Fix]
- TypeError: Field elements must be 2- or 3-tuples, got 1
- ValueError: Expected 2D array, got 1D array instead [Fixed]
- TypeError: ufunc 'isnan' not supported for the input types
- ValueError: columns overlap but no suffix specified [Solved]
- Convert a Row to a Column Header in a Pandas DataFrame
- Shape mismatch: objects cannot be broadcast to a single shape
- Pandas: Create new row for each element in List in DataFrame
- Pandas: Find length of longest String in DataFrame column

